Oniro - The web platform for drug discovery
Want to download, evaluate or know more about ONIRO?
Please refer to Mass
Analytica
Oniro is a web container that encapsulates a number of Molecular Discovery solutions for structure elucidation, in-silico predictions, spectral database, search and reporting that could be used in multiple fields. The platform is distributed together with the WebMetabase, Compound Library, WebChembase and WebQuant applications. It is the way to manage user, settings, import exported files and access to the applications itself.
This application can give access to multiple desktops of server-based applications depending on the apps that the user wants to plug into the system. The different apps can be activated by adding the license line to the code or by setting the connection in the user settings.
This web application controls:
- The licensing for the different applications; individual product licenses have to be requested as usual from this page
- The user access to the different applications as well as the accessibility to the different functionalities inside the applications
- Definition of the workgoups
- Application Settings
- Import experiments from other Oniro installations
The entire Oniro system and databases are designed to be able to interpret, analyze and save Mass Spectra data. Also the system can be used to report and design new compounds in drug discovery and development considering data from multiple sources (vendors, acquisition, experiments, deparments, etc..).
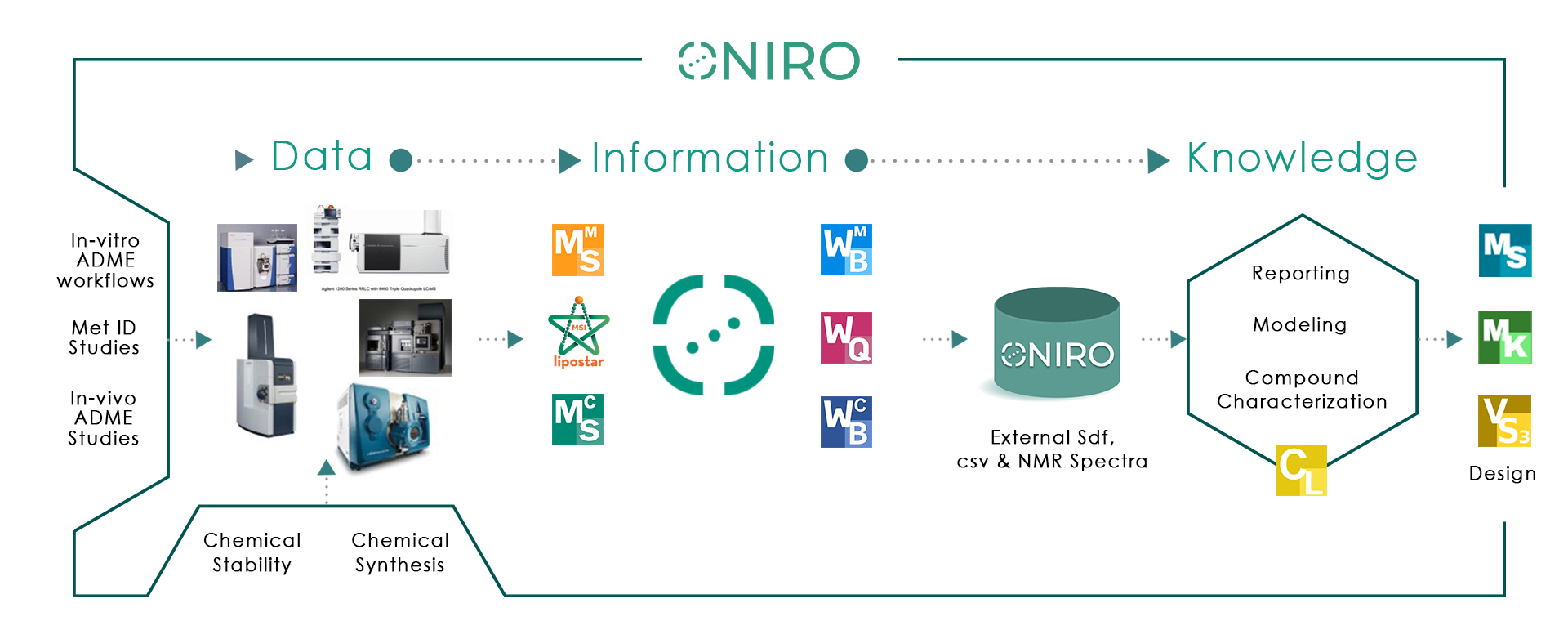
Version 1.3.0 is a major release that introduce many new features in the system, like:
WebMetabase:
- Data process optimization: Common raw data files are converted only once.
- Data enrichment: The New MassMetaSite data values parsed (substrate area %, Met. Description, Iso cluster SIM, Iso SIM)
- Macromolecule:
- Support of new visualization of macromolecule structures with monomers and expanded monomers in Macromolecule experiments (requires MassMetaSite 4.2.3) and updated monomer database.
- Monomer database extension
- Updated default monomers database and introduced new types (like oligos).
- Improved the monomers management system, now the user will be able to organize them in folders and can import custom ones from external JSON files.
WebChembase:
- New Module: Chemical monitoring workflow for MassChemSite experiments.
- Process samples for finding of compounds in complex matrix: pesticides in food, contaminants in the environment, etc.
Compound Library:
- Modeling tool:
- Model monitoring and update system: The new data for existing model can be used to monitor the performance of the model and if needed to automatically update it with the new information.
- Bond breakage modeling and predictions: Ability to predict the most probable fragmentation of a new molecule.
- Model predictions available in:
- WebMetabase, WebChembase and WebQuant experiments.
- in-silico designer.
- MS Spectra tools:
- Import MS/MSMS spectrum from SDF or JSON files.
- Automatic spectrum association with the fragments of a compound in the library.
- External tools:
- Custom MoKa models for Pka prediction
